Spectral mixing matrix normalization
- olivertburton

- Sep 22
- 3 min read
In this post, I talked about the spectral mixing matrix, which is the combination of spectral signatures of our fluorophores that guide our unmixing. I raised the point that we normalize the spectral signatures prior to unmixing to avoid scaling the output unmixed data inversely. In a recent discussion with Sofie Van Gassen, she asked whether the normalization was to the peak emission or to the sum of the emission values across all detectors. Since this is the sort of trivial stuff that I find interesting, let's have a look at what this means.
TLDR: all manufacturers appear to be using peak-based normalization for unmixing.
Here we have some data from human PBMCs run on the Cytek Aurora and unmixed in SpectroFlo (OLS unmixing). I'm reading the data into R so we can plot everything the same way.

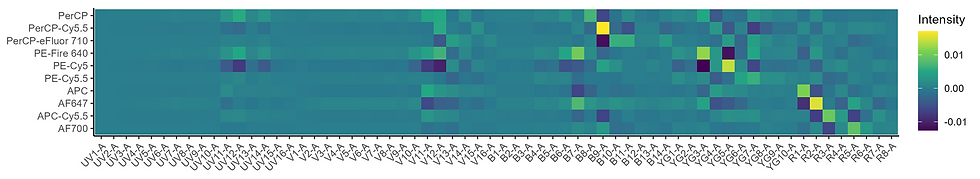
If we normalize each fluorophore to the peak, we get a spectral mixing matrix that looks like this:

If instead, we normalize to the sum of signals across all detectors, essentially area under the curve, we get this:

Notice how the "clean" fluorophores like Spark UV 387 and PerCP-Fire 806 look pretty similar, while "dirty" stuff like BV510 and autofluorescence are much more washed out now.
In this post, we saw that the output unmixed data scale inversely to the spectral mixing matrix values. So, we can expect this area-under-the-curve (AUC) normalization to emphasize fluorophores with signals across more detectors. Why would we want that? Well, when I started doing spectral flow, I'd assumed, naively, that the math would be done that way. One advantage of having extra detectors is that it should allow you to integrate (sum up) signals from all the detectors to build a composite signal for the fluorophore across the entire spectrum. This increases sensitivity for dim, messy fluorophores. As it turns out, that's not what's being done.
If we unmix the raw mixed data using OLS in R, we can compare how it looks with peak-based normalization or AUC normalization. Here's unmixing using OLS with the spectral mixing matrix normalized to the peak:

Looks pretty similar. There are some minor differences, which are down to how the spectral signatures are being pulled from the single-stained controls.
And with the AUC normalization:

Pretty different.
Clearly, we're seeing that amplification of the dim, messy fluorophore values. Both BV510 and BV605 change quite a bit depending on the normalization method.
We're also getting what appears to be more spread in the data. This is a little trickier to determine because the data are now effectively on a different scale, and, being more spread out when plotted on the same scale, the density coloring is also different. It's possible the stain indices could be the same.
In support of there being something going on with spread, we can look at the cosine similarity matrix and the related hotspot matrix.




The values in these matrices are identical. They're based on comparisons using the cosine of the vectors, which is magnitude-independent. That is to say, the cosine doesn't care if there's more or less, just which direction it's heading in.
What is different is the mixing matrix condition number. This tells us something about how problematic the unmixing can be when using least-squares approaches with this spectral mixing matrix. It's higher with the AUC-based normalization. I'd suggest that this makes sense. If we're effectively amplifying the signals from the messier fluorophores, we're going to get more spread in the unmixed data than the other way around, where we favor the cleaner signals. In the case of this staining set, we've got BV510 and very similar autofluorescence signature (cosine 0.91), both of which are greatly affected by the AUC normalization.



Yeah, that doesn't look good. Interesting how we can have such bad unmixing spread with a spreading inflation factor (SIF) of less than 3 between these two.
Anyway, the unmixings for the Cytek Aurora, the Sony ID7000 and the BD FACSDiscover S8 & A8 can all replicated using peak-based normalization of the spectral mixing matrix.







Comments